I recently moved from Kyoto to Paris to do my second postdoc. In the process of flat hunting in Paris, which has been very hard because of the rising housing cost, I walked around some different parts of Paris. One impression I had is Paris is small. Once you walk through the city, you will find this quite easily.
My intuition told me that its size is almost close to the size of Kyoto, which is a medium-sized city in Japan. But, is it true? I found a website comparing London and Paris, and it says London is larger than Paris, but no comparison between Paris and Japanese cities exists.
So, I just checked by superimposing maps of the two cities. I used an R package called “ggmap”, a very nice geographic visualization package built based on ggplot2.
I first downloaded maps of Paris and Kyoto in 15km squares surrounding city centers. (I chose the Cite island for Paris and Shijo-Karasuma for Kyoto as centers). This process was a bit tricky because maps are defined by longitudinal and latitudinal range. The identical longitudinal ranges do not represent the same distance when the cities’ latitudes are different. I used the “great circule distance” equation to adjust the effect of latitude on the longitudinal distance and and find appropriate ranges.
R <- 6371
#convert distance to longitudinal range given a latitude
delta.lambda <- function(d, phi=0.0) {
2*asin(sin(d/R/2)/cos(phi))
}
#create longitudinal and latitudinal ranges which define a suquare with length d
map.range <- function(d, lat=0.0, lon=0.0){
dphi <- d/R*360/2/pi
dlambd <- delta.lambda(d, phi=lat*2*pi/360)*360/2/pi
lonrange <- lon + c(-dlambd/2, dlambd/2)
latrange <- lat + c(-dphi/2, dphi/2)
rang <- c(lonrange, latrange)
names(rang) <- c("left", "right", "bottom", "top")
return (rang)
}
In the above code, the map.range function returns longitudinal and latitudinal range which covers a square with its side length = d km.
Once you can define a range to draw, downloading and plotting maps is simple thanks to ggmap functions.
d <-15 #15kmx15km map
z <- 12 #Zoom level
kyoto_center <- c(35.0, 135.76)
kyoto <- map.range(d, lat=kyoto_center[1], lon=kyoto_center[2])
kyoto_map <- get_stamenmap(kyoto, zoom=z, maptype="toner")
ggmap(kyoto_map)
paris_center <- c(48.855, 2.34)
paris <- map.range(d, lat=paris_center[1], lon=paris_center[2])
paris_map <- get_stamenmap(paris, zoom=z, maptype="toner")
ggmap(paris_map)
The “get_stamenmap” function downloads a map from Stamen Maps. If you like Google map, use “get_googlemap” function to download from Google map.
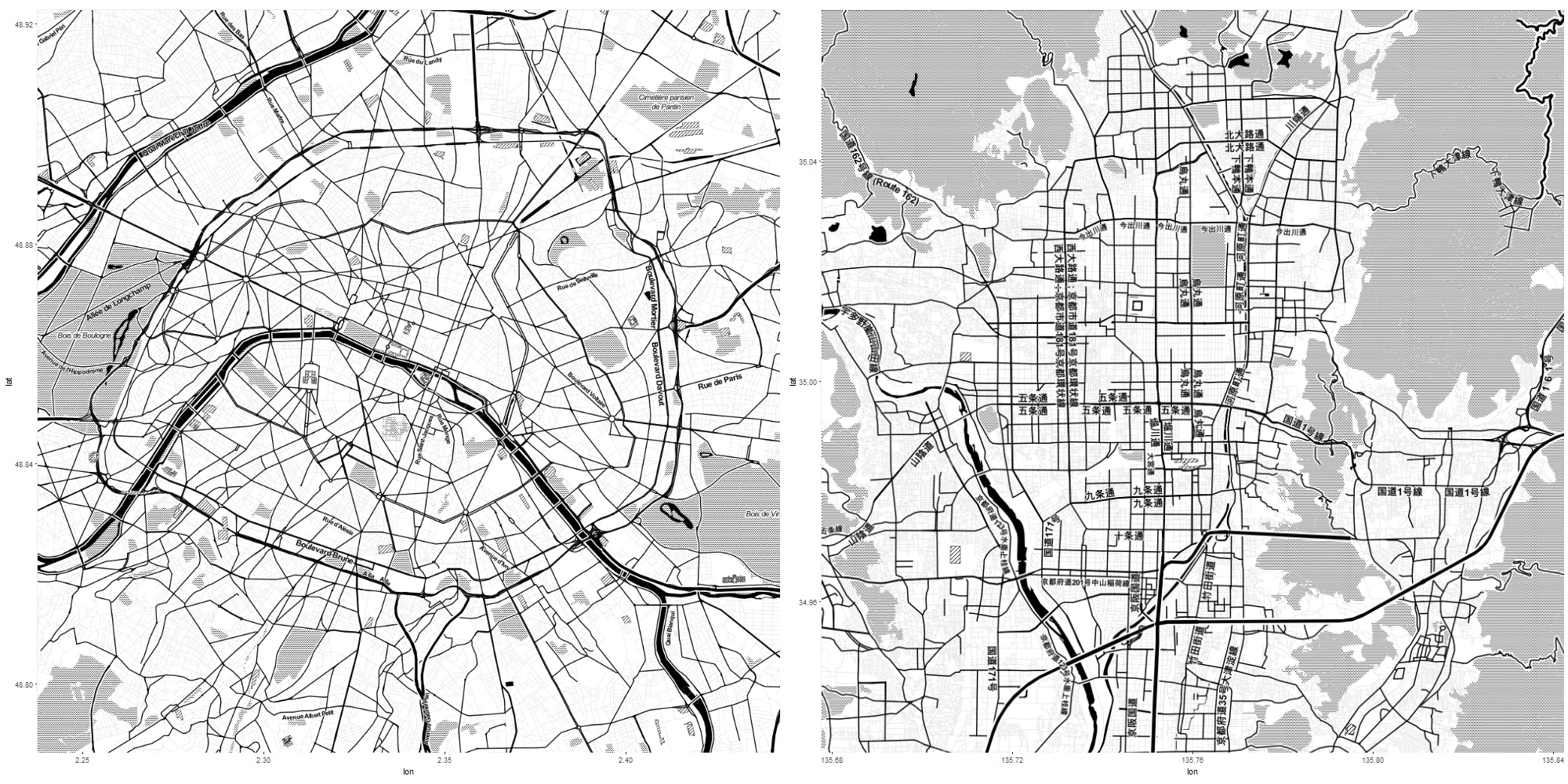
The two maps plotted side by side tells me a lot. For examplem the Peripherique motor way, which defines Paris, looks quite similar size as the city of Kyoto surrounded by mountains.
Superimposing two maps was another tricky point. The ggmap has two ways for superimposition of graphics, “inset_map” and “inset_raster”. It seems “inset_raster” is an appropriate function because it allows me to set coordinates for regions you superimpose graphics. One problem is that it does not allow me to change alpha value. I just added alpha values to the raster graphics of the map and plot it using the inset_raster.
translucent.raster <- function(map){
tmp <- as.matrix(map)
tmp <- gsub("$", "55", tmp)
tmp <- gsub("FFFFFF", "0000FF", tmp)
ras <- as.raster(tmp)
attributes(ras) <- attributes(map)
ras
}
ggmap(paris_map) + inset_raster(translucent.raster(kyoto_map), xmin=paris["left"], ymin=paris["bottom"], xmax=paris["right"], ymax=paris["top"])

This is a bit awkward way, but the result looks OK. Now, I can compare landmarks more precisely.
The Golden and Silver temples (indicated by small black arrows), which define north-west and north-east edges of Kyoto area are almost on the Peripherique. This means walking from the Cite island to the Peripherique is just like walking from Shijo-Karasuma to the Silver temple. Probably, this distance similarity made me feel that the two cities are in similar size.
Of course, Paris has much larger suburban area, and the size of Paris metropolitan area is much larger than Kyoto. But, my intuition looks more or less correct.